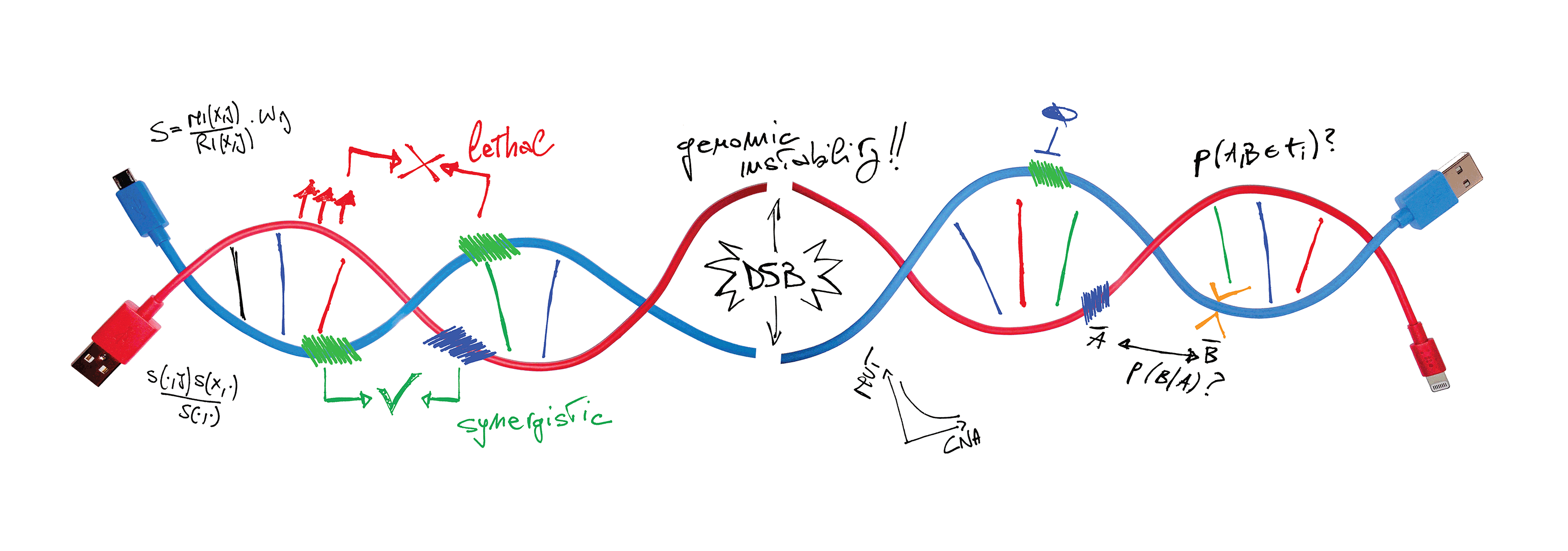


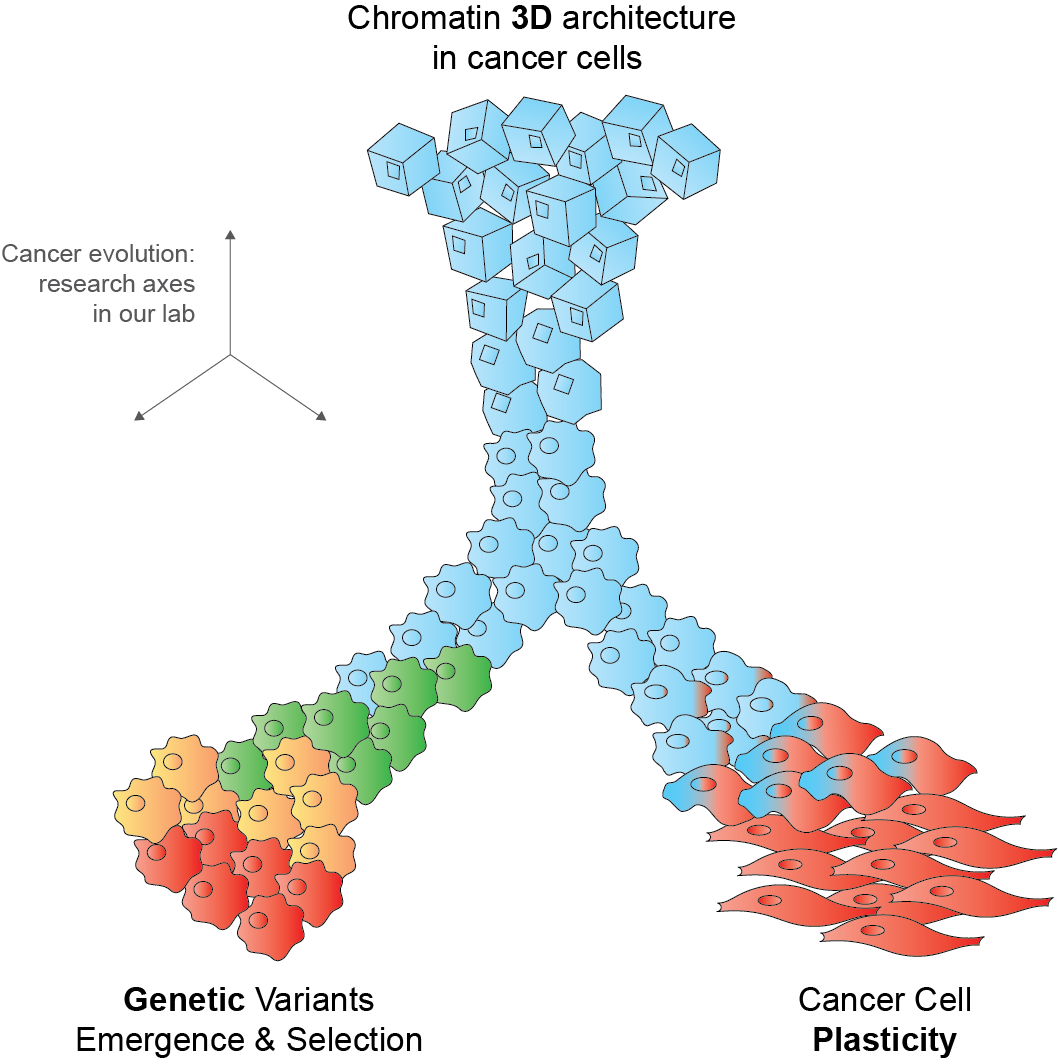
Cancer emerges through the occurrence and selection of molecular alterations. We aim at understanding factors that favor or veto selection of specific alterations, a.k.a. evolutionary dependencies (EDs). In particular, we focus on concurrent or mutually exclusive selection of genetic alteations and whether these EDs can inform response to therapy.
Cancer cells can change their phenotype or even their identity without modifying their genetic code. We are interested in understanding epigenetic and transcriptional reprogramming programs that underlie cancer cell plasticity. We study features of plastic reprogramming among different patients and within individual tumors, using cutting edge single cell and spatial -omics technologies.
A key paradigm in biology is that structure determines function. Whether and to what extent this holds true for chromatin 3D architectures remains an open question. Here, we study chromatin structural changes in response to cancer genetic variants and epigenetic reprogramming. Our goal is to decipher chromatin plasticity and determine how it is hijacked in and/or it influences tumor phenotypes.
Currently open positions:
